Toy Example - Parametric prior#
import tensorflow_probability as tfp
import matplotlib.pyplot as plt
import tensorflow as tf
import seaborn as sns
import pandas as pd
import numpy as np
import elicit as el
import copy
from elicit.extras import utils
tfd = tfp.distributions
The Model#
Generative model#
Implementation#
Generative model#
class ToyModel:
def __call__(self, prior_samples, design_matrix):
# linear predictor
epred=tf.matmul(prior_samples[:,:,:-1], design_matrix,
transpose_b=True)
# data-generating model
likelihood = tfd.Normal(
loc=epred, scale=tf.expand_dims(prior_samples[:, :, -1], -1)
)
# data-generating model
likelihood = tfd.Normal(
loc=epred, scale=tf.expand_dims(prior_samples[:, :, -1], -1)
)
# prior predictive distribution (=height)
ypred = likelihood.sample()
# selected observations
y_X0, y_X1, y_X2 = (ypred[:,:,0], ypred[:,:,1], ypred[:,:,2])
# R2
var_ypred = tf.math.reduce_variance(ypred, -1)
var_epred = tf.math.reduce_variance(epred, -1)
R2 = tf.math.divide(var_ypred, tf.add(var_ypred, var_epred))
return dict(
y_X0=y_X0, y_X1=y_X1, y_X2=y_X2,
R2=R2
)
Define design matrix#
# create a predictor ranging from 1 to 200
# standardize predictor
# select the 25th (X0), 50th (X1), and 75th (X2) quantile of the std. predictor for querying the expert
def X_design(N, quantiles):
X = tf.cast(np.arange(N), tf.float32)
X_std = (X-tf.reduce_mean(X))/tf.math.reduce_std(X)
X_sel = tfp.stats.percentile(X_std, quantiles)
X_design = tf.stack([tf.ones(X_sel.shape),X_sel], -1)
return X_design
X_design(N=200, quantiles=[25,50,75])
<tf.Tensor: shape=(3, 2), dtype=float32, numpy=
array([[ 1. , -0.85737586],
[ 1. , 0.00866036],
[ 1. , 0.85737586]], dtype=float32)>
Model input for elicit method#
# specify the model
model=el.model(
obj=ToyModel,
design_matrix=X_design(N=200, quantiles=[25,50,75])
)
Model parameters#
intercept with normal prior \(\beta_0\)
slope with normal prior \(\beta_1\)
random noise with halfnormal prior \(\sigma\)
To be learned hyperparameters \( \lambda = (\mu_0, \sigma_0, \mu_1, \sigma_1, \sigma_2) \)
scale parameters (\(\sigma_0, \sigma_1, \sigma_2\)) are constrained to be positive
Parameter input for elicit method#
parameters=[
el.parameter(
name="beta0",
family=tfd.Normal,
hyperparams=dict(
loc=el.hyper("mu0"),
scale=el.hyper("sigma0", lower=0)
)
),
el.parameter(
name="beta1",
family=tfd.Normal,
hyperparams=dict(
loc=el.hyper("mu1"),
scale=el.hyper("sigma1", lower=0)
)
),
el.parameter(
name="sigma",
family=tfd.HalfNormal,
hyperparams=dict(
scale=el.hyper("sigma2", lower=0)
)
),
]
Target quantities and elicitation techniques#
Target quantities
query expert regarding prior predictions \(y \mid X_{i}\) with \(i\) being the 25th, 50th, and 75th quantile of the predictor.
Elicitation technique
query each observation using quantile-based elicitation using \(Q_p(y \mid X)\) for \(p=5, 25, 50, 75, 95\)
Importance of elicited statistics in loss
all elicited statistics should have equal importance (weight=1.0)
for computing the discrepancy between expert-elicited statistics and model simulations with use the Maximum Mean Discrepancy with Energy kernel
Targets input for elicit method#
targets=[
el.target(
name="y_X0",
query=el.queries.quantiles((.05, .25, .50, .75, .95)),
loss=el.losses.MMD2(kernel="energy"),
weight=1.0
),
el.target(
name="y_X1",
query=el.queries.quantiles((.05, .25, .50, .75, .95)),
loss=el.losses.MMD2(kernel="energy"),
weight=1.0
),
el.target(
name="y_X2",
query=el.queries.quantiles((.05, .25, .50, .75, .95)),
loss=el.losses.MMD2(kernel="energy"),
weight=1.0
),
el.target(
name="R2",
query=el.queries.quantiles((.05, .25, .50, .75, .95)),
loss=el.losses.MMD2(kernel="energy"),
weight=1.0
)
]
Expert elicitation#
instead of querying a “real” expert, we define a ground truth (i.e., oracle) and simulate the oracle-elicited statistics
Expert input for elicit method (here: oracle)#
# specify ground truth
ground_truth = {
"beta0": tfd.Normal(loc=5, scale=1),
"beta1": tfd.Normal(loc=2, scale=2),
"sigma": tfd.HalfNormal(scale=5.0),
}
# define oracle
expert=el.expert.simulator(
ground_truth = ground_truth,
num_samples = 10_000
)
Training: Learn prior distributions based on expert data#
All inputs for elicit method
eliobj = el.Elicit(
model=model,
parameters=parameters,
targets=targets,
expert=expert,
optimizer=el.optimizer(
optimizer=tf.keras.optimizers.Adam,
learning_rate=tf.keras.optimizers.schedules.CosineDecay(0.1,400),
clipnorm=1.0
),
trainer=el.trainer(
method="parametric_prior",
seed=1,
epochs=400
),
initializer=el.initializer(
method="random",
loss_quantile=0,
iterations=32,
distribution=el.initialization.uniform(
radius=3.,
mean=3.
)
)
)
Run multiple chains in parallel
eliobj.fit(parallel=el.utils.parallel(chains=4))
Save eliobj
# save eliobj
eliobj.save(name="m1", overwrite=True)
saved in: ./results/parametric_prior/m1_1.pkl
Results#
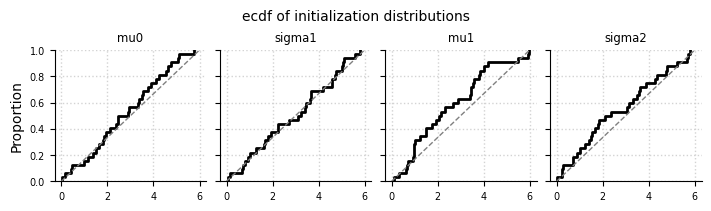
Initialization of hyperparameters#
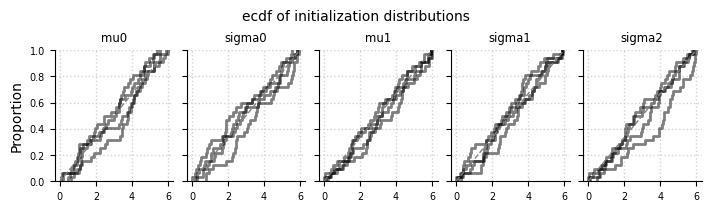
el.plots.initialization(eliobj, cols=5, figsize=(7,2))
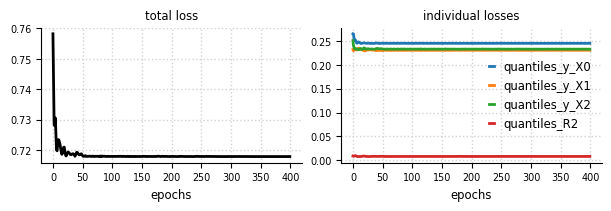
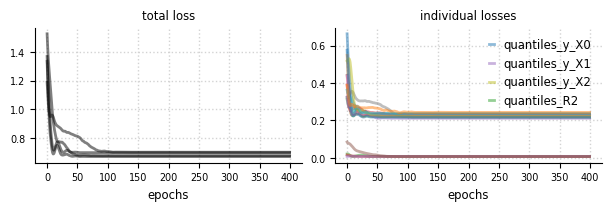
Convergence#
el.plots.loss(eliobj, figsize=(6,2))
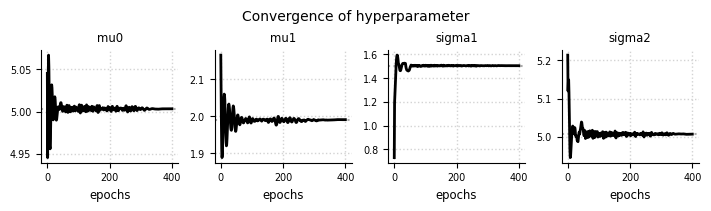
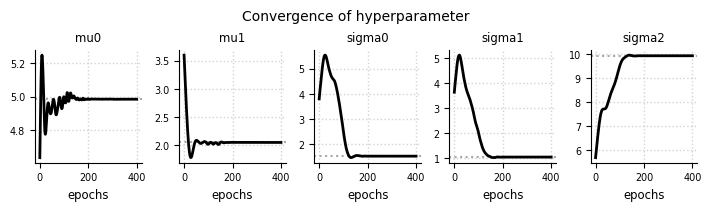
el.plots.hyperparameter(eliobj, cols=5, figsize=(7,2))
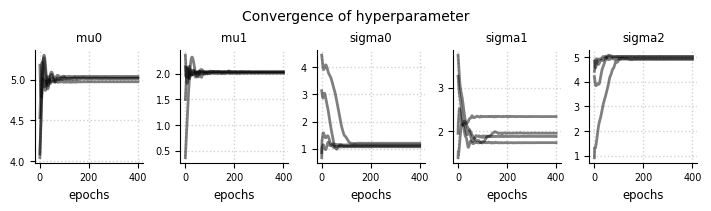
Expert expectations#
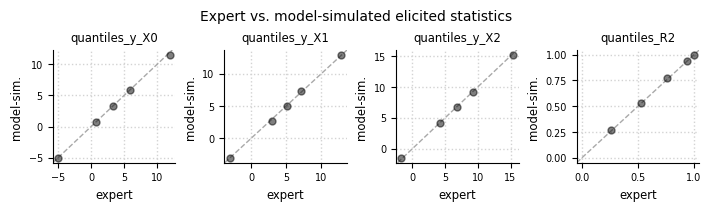
el.plots.elicits(eliobj, cols=4, figsize=(7,2))
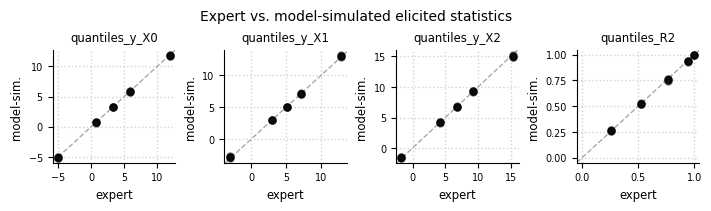
Learned priors#
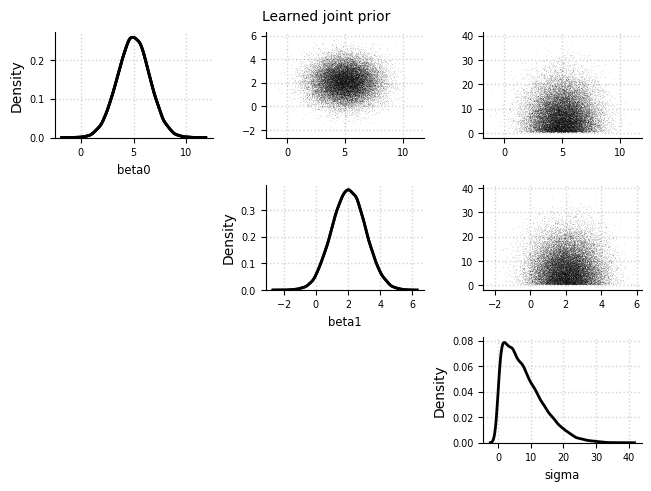
Joint prior#
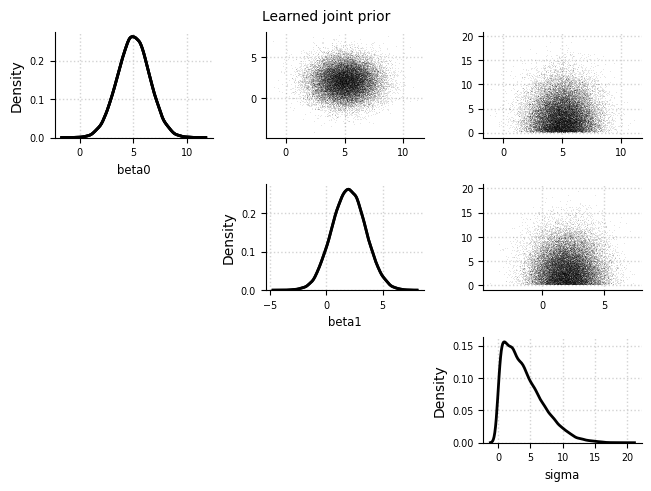
el.plots.prior_joint(eliobj, idx=0)
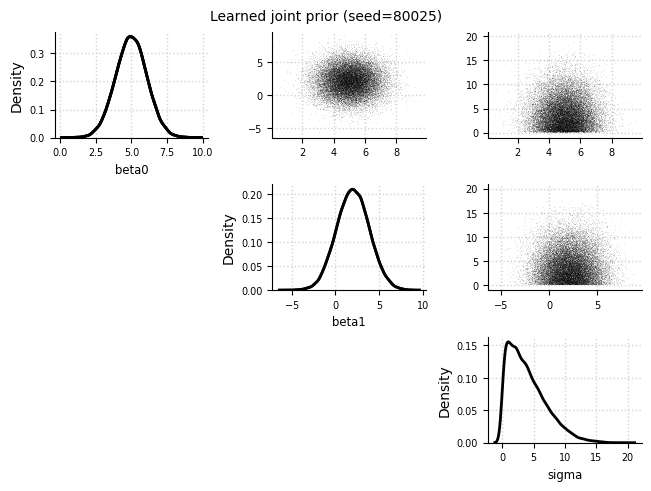
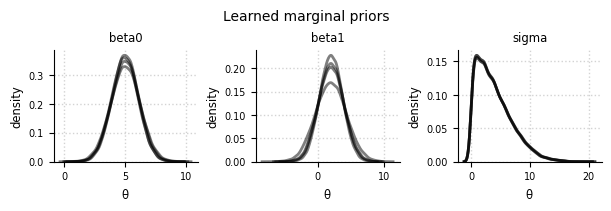
Marginal priors#
el.plots.prior_marginals(eliobj, figsize=(6,2))
INFO: Reset cols=3 (number of elicited statistics)
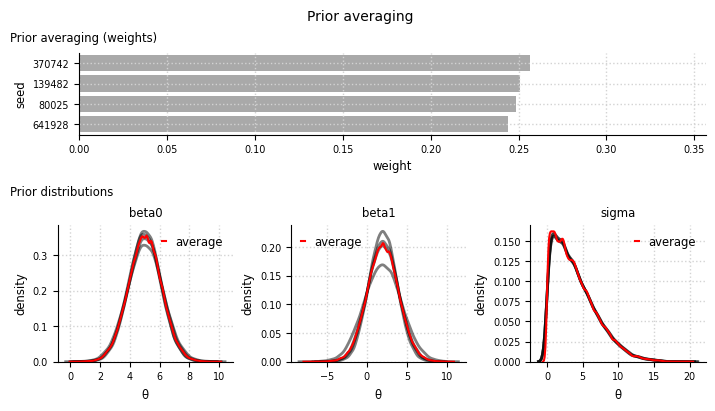
Model averaging#
el.plots.prior_averaging(eliobj, cols=3, figsize=(7,4), height_ratio=[1,1.5])
Add-on: Use expert data as input#
# create a copy of eliobj
eliobj_dat = copy.deepcopy(eliobj)
# use dictionary of elicited expert data (instead of simulating data)
expert_dat = {
"quantiles_y_X0": [-5.0, 0.7, 3.3, 5.9, 11.9],
"quantiles_y_X1": [-3.1, 3.0, 5.0, 7.1, 12.9],
"quantiles_y_X2": [-1.7, 4.2, 6.8, 9.3, 15.4],
"quantiles_R2": [0.26, 0.53, 0.76, 0.94, 0.99]
}
# update expert in eliobj
eliobj_dat.update(expert=el.expert.data(dat = expert_dat))
# refit the model and run again once
eliobj_dat.fit()
INFO: Results have been reset.
Initialization
100%|██████████| 32/32 [00:04<00:00, 7.91it/s]
Training
100%|██████████| 400/400 [02:06<00:00, 3.17it/s]
Results#
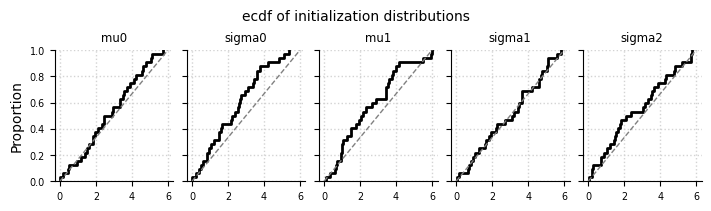
Initialization of hyperparameters#
el.plots.initialization(eliobj_dat, cols=5, figsize=(7,2))
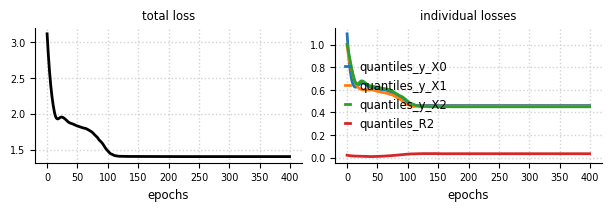
Convergence#
el.plots.loss(eliobj_dat, figsize=(6,2))
el.plots.hyperparameter(eliobj_dat, cols=5, figsize=(7,2))
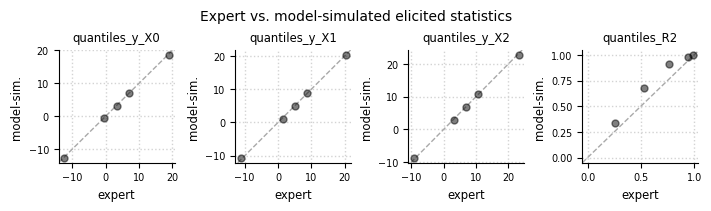
Expert expectations#
el.plots.elicits(eliobj_dat, cols=4, figsize=(7,2))
Learned priors#
el.plots.prior_joint(eliobj_dat)